

| | |||||||||||||||||
| |||||||||||||||||
Table of Contents
- 1. General Questions
- 1.1 What is ModeRNA?
- 1.2 How to cite ModeRNA?
- 1.3 How much does it cost?
- 1.4 What system does ModeRNA work on?
- 1.5 How long does it take to build a model?
- 1.6 What language is ModeRNA written in?
- 1.7 How to get help with the program?
- 1.8 What command-line options are available
- 1.9 How to use ModeRNA commands interactively?
- 1.10 How to write an input script?
- 2. Building Models with ModeRNA
- 2.1 What do I need for automatic modeling?
- 2.2 How to prepare the template structure?
- 2.3 How to prepare the alignment?
- 2.4 How to build a model from a template and alignment?
- 2.5 How to write a model to a PDB file?
- 2.6 Can I build models with multiple chains?
- 2.7 Can I build models from multiple templates?
- 2.8 How to fix breaks in the backbone?
- 3. Modified Nucleotides
- 4. Loop/indel Modeling
- 5. Analyzing Structures
- 6. Refinement
- 7. Advanced use of ModeRNA
- 8. Tips & Tricks
1. General Questions
1.1 What is ModeRNA?
ModeRNA is a program for 3D modeling of RNA structures. It uses a homology modeling approach, taking an alignment and a template structure as an input. It contains many functions supporting analysis and manipulation of many nucleic acid structures.
1.2 How to cite ModeRNA?
ModeRNA: a tool for comparative modeling of RNA 3D structure.
Rother M, Rother K, Puton T, Bujnicki JM.
Nucleic Acids Res. 2011 Feb 7. [Epub ahead of print]
1.3 How much does it cost?
ModeRNA is free of charge for academic and commercial users. The code is published under the conditions of the Python License.
1.4 What system does ModeRNA work on?
ModeRNA has been tested on both Linux and Windows successfully. Both require Python and a couple of libraries to be installed. Some extra functions, like the model optimization with MMTK will not run on Windows computers. We have not tested whether ModeRNA runs on a Mac, sorry.
1.5 How long does it take to build a model?
Preparation of the input sequence alignment and template structure involves manual work and can be time consuming. For complicated cases, an input script needs to be written (or copied from this page). Once everything is prepared, the modeling of a tRNA-sized structure takes about half a minute. If the sequence alignments and templates are prepared, ModeRNA can automatically produce 10,000 tRNA models over a weekend on an average PC.
1.6 What language is ModeRNA written in?
The code behind ModeRNA is 100% Python. It uses BioPython, NumPy, and PyCogent for particular tasks.
1.7 How to get help with the program?
First, try to read the Tutorial. If you need help with the command-line interface, try:
python moderna.py -h
1.8 What command-line options are available
ModeRNA has a straightforward command-line interface. To use it you need to open a console
and enter a command according to this pattern:
python moderna.py <option1> <argument1> <option2> <argument2> ...
Available options:
| option | argument | name | comments |
|---|---|---|---|
| -h | none | help | Prints available options. |
| -a | .fasta file | alignment | Reads an alignment from the given fasta file. |
| -t | .pdb file | template | Reads a template structure from the given file. |
| -e | - | examine | Checks the template for irregularities. |
| -l | - | clean | Removes irregularities (water, ions, amino acids, nonstandard atom names) from a PDB structure. |
| -c | chain identifier | chain ID | Specifies the chain read from the structure given by -s or -t (default is A). |
| -o | .pdb file | Writes the model structure to the given file (by default ModeRNA.out.pdb). | |
| -s | .pdb file | structure | Reads the structure in the given file for adding modifications. |
| -m | abbreviation | modification | Adds a nucleotide modification (abbreviated like m2G, mnm5U, Y) |
| -p | residue number | position | Position at which a modification is to be added. |
1.9 How to use ModeRNA commands interactively?
ModeRNA can be used from the Python interactive shell (similar to BioPython). Just open a console and type python.
You should see a prompt (>>>). Activate ModeRNA by typing:
>>> from moderna import *
1.10 How to write an input script?
An input script is a normal text file containing ModeRNA commands, executed by the Python interpreter. The first command always needs to be the following:
from moderna import *
python <my_script>On Windows, you need the proper path settings, so that the system finds the Python.exe file.
2. Building Models with ModeRNA
2.1 What do I need for automatic modeling?
For building an RNA model automatically, you need a template structure and an alignment file. The template should cover all sections of the target sequence, leaving no gaps larger than 15 bases in the alignment.
[ example template structure ]
[ example alignment ]
Caution: Casual modelers must be warned that for large RNA molecules with complex structures, the development of a good alignment may require laborious manual preparation of the input data based on previous expertise of the respective RNA family.
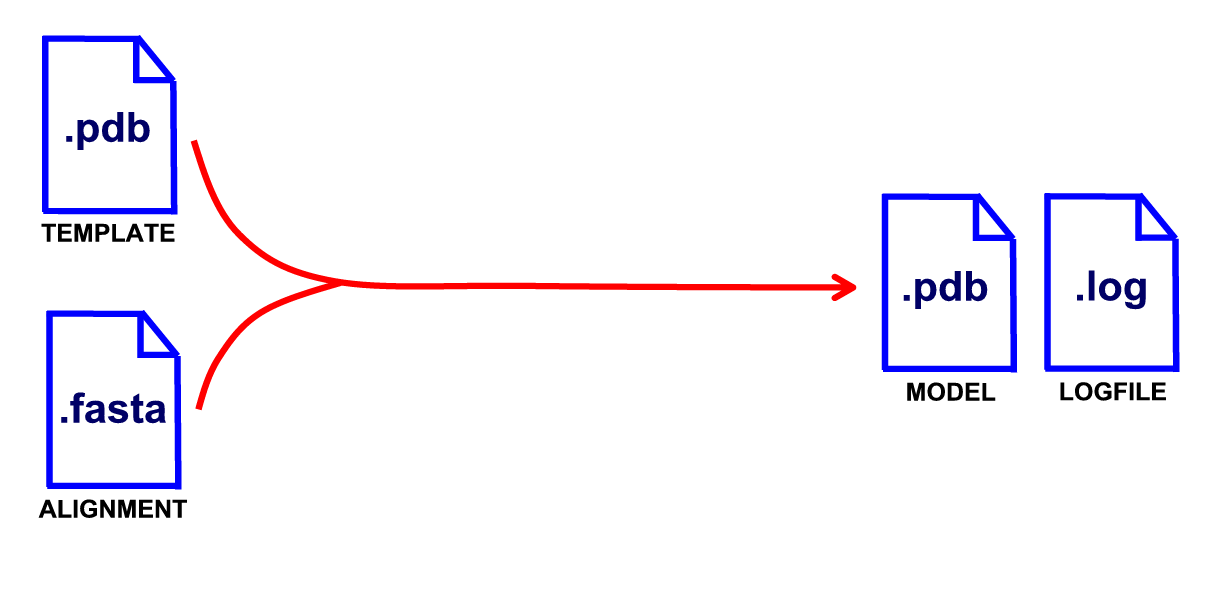
2.2 How to prepare the template structure?
Atoms in the RNA backbone and ribose units need to comply to the remediated standard names:
P, OP1, OP2, C1', C2', O2', C3', etc.. The atoms need to be complete.
In the bases, at least the N1 or N9 atom (the first after the glycosidic bond) must be present.
It does not matter, how modified nucleotides are named, or whether they are HETATM or ATOM records.
Any remediated PDB file complies with these requirements. See the according section in Concepts for details.
2.3 How to prepare the alignment?
For producing reasonable models, the second sequence in the alignment needs to be identical to the one in the template structure. This example shows how to get the sequence from a given template structure in order to insert it to an alignment.
# write the template sequence to screen
t = load_template('1QF6_B_tRNA.pdb','B')
print get_sequence(t)
# check whether template and alignment have the same sequence
a = load_alignment('aln_1QF6_E_coli_1C0A.fasta')
print match_template_with_alignment(t,a)
> target sequence GDCGGD--DAGAAUACCUGCCUQUCACGCAGGG-----CGGGTPCGAGU > template sequence GDCGGDCUDAGAAUA----CCUQUCACGCA--GG7UCGCGGGTPCGAGUTo have ModeRNA handle gaps automatically, they may not be closer than two bases to each other. In the example, for the first two gaps a linker will be inserted, but not for the second two.
2.4 How to build a model from a template and alignment?
With a prepared template structure and alignment file, a model can be created from the Python shell with a few commands:
t = load_template('1QF6_B_tRNA.pdb', 'B')
a = load_alignment('aln_1QF6_E_coli_1C0A.fasta')
m = create_model(t,a)
m.write_pdb_file('my_model.pdb')
2.5 How to write a model to a PDB file?
Every model can be written as a PDB file. The residues are saved in the order of their respective numbers.
write_model(m)
write_model(m, 'output.pdb')
write_model(m, 'output.pdb', 'log.txt')
2.6 Can I build models with multiple chains?
At present, ModeRNA does not handle more than one chain in the same model. It is nevertheless possible to merge residues from two chains to one model. The model will then contain two regions with different numbers, and the delimiter between the chains can be seen in the sequence as a '_' symbol. See 7.4 and 7.5 for details.
2.7 Can I build models from multiple templates?
ModeRNA can combine multiple templates to build one model. In such scenarios, one template serves
as the 'core' structure, and other templates are attached to it. The attachment is done by defining one or two
linker residues (one, when attaching to the 5' or 3' end, two when attaching the second template within the core.
After combining two templates, the backbone may be interrupted, and thus the fix_backbone()
function should be used afterwards.
See 2.8, 7.4, and 7.5 for details.
2.8 How to fix breaks in the backbone?
Both template structures from the PDB and models built with ModeRNA sometimes contain chemically unreasonable
interatomic distances along the backbone. These can be recognized and fixed by the fix_backbone()
function.
m = load_model('broken_bb.pdb','A')
print m.get_sequence()
# check and fix the entire model
fix_backbone(m)
print m.get_sequence()
# check and fix the connection between residues 4 and 5
fix_backbone(m, '4', '5')
3. Modified Nucleotides
3.1 How to add a single modification?
If a model exists, but a single modification needs to be introduced, this can be achieved from the command line by:
moderna.py -s my_structure -m m2G -p 63to introduce a 2-methyl-guanosine in position 63.
From a script, the same operation looks like:
# add a 2-methyl-Guanosine in position 63.
m = load_model('1QF6_B_tRNA.pdb', 'B')
add_modification(m['63'], 'm2G')
3.2 Which modifications are present in my structure?
Given a template or a model, it is possible to find all modifications in a structure by:
# find modifications in any structure file
t = load_template('1QF6_B_tRNA.pdb', 'B')
print find_modifications(t)
‘634’: <Residue Q het=H_Q resseq=634 icode= >}
3.3 How to remove modified nucleotides?
If neccesary, all modifications can be removed from a structure by:
remove_all_modifications(m)
3.4 How to obtain a full list of modified nucleotides?
A full list of nucleotides, and their abbreviations can be found in the MODOMICS database. With the exception of very few recently added entries, all of them are available from within ModeRNA.
4. Loop/Indel Modeling
4.1 How to insert a single loop?
The LIR loop library of ModeRNA can be searched for suitable linkers between two nucleotides. An internal scoring is done to select the best suitable candidate. To find the best fitting fragment and add it to your model, you need to use:
# prepare a model
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_some_residues(t['31':'35']+t['38':'42'],m)
# Find the best fitting fragment for the missing two
apply_indel(m, '35', '38', Sequence('CA'))
4.2 How can I select manually from several loop candidates?
If the result of the automatic loop insertion is not satisfactory, you can find a set of loop candidates to PDB files in a given directory and inspect them manually (by default 20):
MISSING EXAMPLE: write_loop_candidates
To insert e.g. the 3rd candidate, use:# the index starts counting at zero.
insert_fragment(m, candidates[2])
4.3 How to add my own structural fragment in a certain place?
You can also close gaps in the model by adding custom pieces of structure. To do so, you need to provide a PDB file with the fragment plus one extra residue on each side, which is used for superimposition with the model. E.g. for adding a fragment two residues long, the file should contain four residues [example file]:
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_some_residues(t['31':'35']+t['38':'42'],m)
# inserts the fragment between the indicated residues.
f = create_fragment('my_fragment.pdb', m['35'], m['38'], 'B')
insert_fragment(m,f)
4.4 How does the search for loop candidates work?
The loop search procedure in ModeRNA tries to find the best fitting
loop from a library of 88000 RNA fragments. The search is done in two
steps: A fast one based on geometrical parameters, and a slower one
based on the exact fit of a smaller set of loop candidatas. In the
first step, the similarity of seven parameters derived from the
residues at both ends of the fragment is compared to the same
parameters of the residues in the model, between which a linker is to
be found. These parameters involve three distances, two angles, and
three dihedrals (two of which enforce a proper orientation of the
bases). Additionally, the sequence similarity of the linker and the
desired fragment is taken into account. A description of this approach
can be found in (Michalsky E, Goede A, Preissner R. Loops In Proteins (LIP)–a comprehensive loop database for homology modelling. Protein Eng. 2003 Dec;16(12):979-85.).
In the second step, the 20 candidate loops that are most similar
according to these criteria are inserted into the model, superimposing
the P,O5',C5' atoms of the terminal residues at the 5' end, and
C4',C3',O3' at the 3' end. A combined score is calculated, using the
RMSD of the superposition, the number of inter-residue clashes, the
number of hydrogen bonds formed by the loop, and the score used in the
first step. The best ranking candidate is then inserted, replacing the
two terminal residues from the model by the loop candidates'.
When building models automatically, ModeRNA eliminates one extra
residue on each side of a gap to be inserted, to give room for a
reasonable loop closure.
5. Analyzing Structures
5.1 How to analyze the geometry of a structure?
Apart from modeling tasks, ModeRNA is capable of checking whether bond lengths, bond angles, and torsion angles are in a valid range. To measure the geometry of a given file, do:
t = load_template('1QF6_B_tRNA.pdb', 'B')
analyze_geometry(t)
moderna.log file.
The allowed range for bond lengths and angles was determined by running statistics on a the RNADB2005
set of structures. Torsion angles for the RNA backbone were taken from (Murray, Richardson et al. 2008). The
torsion angles of the ribose were measured, and for the χ angle every value is considered valid.
5.2 How to check for interatomic clashes?
With ModeRNA the user can check whether model contains interatomic clashes. The list of clashing residue pairs can be obtained:
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_some_residues(t['31':'35']+t['38':'42'],m)
candidates = find_fragment(m, '35', '38', Sequence('CAG'), 20)
insert_fragment(m, candidates[19])
find_clashes(m)
6. Refinement
6.1 What programs can refine RNA models?
Once ModeRNA has produced a model, it is often necessary to refine the local geometry. MMTK is a program library that performs energy minimization using Conjugate Gradient and others. It uses the AMBER force field. Alternatively, it is also possible to use a different Molecular Dynamics program like NAMD, or forcefield like Charmm. Some commercial packages like HyperChem are also capable of performing energy minimization of RNA.
6.2 How to refine some parts of a model?
The script refine_model_mmtk.py allows to improve sections of the model using
MMTK (it should on $PATH environment variable after installation).
It prepares the structure for input (basically exchanging some residue and atom
names) and starts a run of energy minimization. For a given part, the first and last
residues will be fixed by applying strong distance, angle and torsion restraints. This
way, it is made sure that the optimized part fits into the rest of the model.
Example:
refine_model_mmtk.py -m 1QF6_B_tRNA.pdb -c B -r 50-55 -y 1000\ -o optmized.pdb
6.3 How to refine an entire model?
It is also possible to optimize an entire model structure with the refine_model_mmtk.py script.
This may be necessary, e.g. to resolve conflicts between different parts of the chain.
Optimization takes considerably longer than for a single fragment (several hours for a tRNA-sized structure).
Example:
refine_model_mmtk.py -m 1QF6_B_tRNA.pdb -c B -y 1000 -o optmized.pdb
6.4 How to refine modifications
Modified nucleotides are not part of the standard AMBER forcefield. They are therefore excluded from the refinement by the MMTK script. A set of quantum mechanical parameters for 111 modified nucleotides has been published by (Aduri R et al. J Chem Theory Comput, 2007, 3(4), 1464–1475). They can be included in the AMBER parameter files to allow modifications to be refined in the same way.
7. Advanced use of ModeRNA
7.1 Can ModeRNA also model DNA?
Yes, this is possible in exactly the same way. In the sequence alignment, DNA is marked by lowercase letters.
> target sequence: DNA aaagggcccttt > template sequence: homologous RNA AAAGGGCCCUUU
7.2 How to combine pieces of structure?
Almost any structural fragments can be combined together using ModeRNA. As an example, 3 small pieces of structures (11, 12 and 11-residues long) are connected into one structure. The fragment in the middle is loaded as a model (m), then the fragment which will be added at 5’ end is loaded as a fragment (f1), the same for the fragment which will be added at 3’ end (f2). Finally, the fragments are added:
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_some_residues(t['31':'35']+t['38':'42'],m)
# inserts the fragment between the indicated residues.
f = create_fragment('my_fragment.pdb', m['35'], m['38'], 'B')
insert_fragment(m,f)
7.3 Can I write Python programs using ModeRNA?
Yes, all ModeRNA input scripts are essentially Python programs. Thus, it is straightforward to write Python-ModeRNA hybrid code.
from moderna import *
m = load_model('1QF6_B_tRNA.pdb','B')
for resi in m:
if resi.modified:
print 'Modified base:',resi
7.4 How to merge two chains into one?
Two chains can be merged by copying the residues and adjusting the numbers:
# building a model composed of two strands
strand_a = load_template('dr0005H.pdb', 'A')
strand_b = load_template('dr0005H.pdb', 'B')
m = create_model()
copy_some_residues(strand_a['3':'21'],m)
renumber_chain(strand_b,101)
copy_some_residues(strand_b['101':'111'],m)
[ download example structure ]
7.5 How to combine two template structures?
Multiple templates can be combined by the same mechanism used in modeling linkers. A template is loaded as a fragment, and attached by either one or two residues to the existing structural core. Then, eventual discontinuities in the backbone are detected and fixed automatically:
# load the template for the structural core
m = load_model('1C0A.pdb', 'B')
# add a second template that is attached to the core.
f = create_fragment('hairpin_14nt.pdb', m['631'], m['639'], 'R')
insert_fragment(m,f)
fix_backbone(m)
[ download example structure ] [ download extension ]
8. Tips & Tricks
8.1 How to extract one chain from a structure?
When ModeRNA loads a template structure, it only considers one of the chains (A by default) - everything else is discarded. This function can be used to store nucleic acid chains in separate files for convenience:
m = load_model('1QF6_B_tRNA.pdb', 'B')
write_model(m, 'result.pdb')
8.2 How to copy a part of a template?
An arbitrary fragment of structure can be copied from a template to the model by:
from moderna import *
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_some_residues(t['1':'15'],m)
8.3 How to copy a single residue?
It is also possible to copy only one residue in a similar way:
from moderna import *
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_single_residue(t['37'], m)
8.4 How to exchange bases?
Standard bases can be exchanged by:
from moderna import *
t = load_template('1QF6_B_tRNA.pdb', 'B')
m = create_model()
copy_single_residue(t['37'], m)
exchange_single_base(t['36'], 'C', m)
8.5 How to clean up a template structure?
A template (or any PDB structure) can be analyzed for irregularities and be cleaned up by:
# cleaning up a loaded template:
# removes water, ions, amino acids, and unknown residues
# replaces O1P and O2P in atom names by OP1 and OP2
# replaces * in atom names by '
clean_structure(t)
|
| Copyright© - Adam Mickiewicz University - All rights reserved |